Chemistry, Department of: Faculty Series
Mark A. Griep Publications
Accessibility Remediation
If you are unable to use this item in its current form due to accessibility barriers, you may request remediation through our remediation request form.
Document Type
Article
Date of this Version
6-1-2006
Abstract
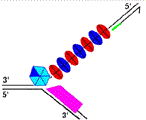
The polymerase chain reaction is one of the most important reactions in molecular biology. Single stranded DNA is copied in a complex series of steps, at the core of which lies the action of the DNA polymerase. At each nucleotide along the template, the polymerase screens the dNTP pool until it finds the complementary dNTP. The insertion of each dNMP is a balance between high fidelity and rapid elongation. In this study the kinetics of the β type polymerase pyrococcus kodakaraensis (KOD) is analyzed. The kinetics is influenced by reaction conditions such as the dNTP pool composition and temperature. In a previous study by Viljoen et al. [2005, A macroscopic kinetic model for DNA polymerase elongation and high-fidelity nucleotide selection. Computational Biology and Chemistry 29, 101–110], a macroscopic kinetics expression of the polymerase chain reaction has been derived. The model contains four parameters that are intrinsic to a specific polymerase. The experiments to measure the temperature- dependence of the parameters for KOD DNA polymerase are reported. The results indicate that the optimal temperature for an equimolar dNTP pool is 72.5 °C and the optimum temperature shifts to lower temperatures when the dNTP pool composition is biased.


Comments
Published in Chemical Engineering Science 61:12 (June 2006), pp. 3885–3892. doi:10.1016/j.ces.2005.12.032 Copyright © 2006 Elsevier Ltd. Used by permission. http://www.sciencedirect.com/science/journal/00092509