Chemistry, Department of: Faculty Series
Robert Powers Publications
Document Type
Article
Date of this Version
September 2004
Abstract
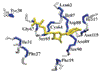
Structural genomics is providing a means to determine the molecular and cellular function for the vast amount of proteins in the Human proteome that lack any explicit experimental information by characterizing the complete range of protein folds (Montelione, 2001). The Northeast Structural Genomics Consortium (NESG; http://www.nesg.org/) is a pilot project funded by the National Institutes of Health Protein Structure Initiative, focusing on proteins from eukaryotic model organisms including humans. The thermophillic archaea Archaeglobus fulgidis AF2095 protein is an example of a protein of unknown biological function targeted for structural analysis by NESG. AF2095 belongs to the Pfam family PF01981 – UPF0099, protein domain family of unknown function that has been found in yeast, archaebacteria and eubacteria. AF2095 has been assigned to NESG Cluster ID:17431, a set of fourteen proteins with high (>~30%) sequence identity with human, Drosophila, Caenorhabditis elegans, Arabidopsis, yeast, archaeal and eubacterial origin (Liu, 2004). A total of fifty-six proteins are identified when the analysis is expanded to include all available genomes, where determining the NMR solution structure of AF2095 can be leveraged to infer 3D structural information for these proteins. Here we report the near complete 1H, 15N, and 13C NMR as signments and secondary structure of AF2095. These data provide a basis for determining the solution structure of AF2095, for further investigation of the func tion of this protein and for providing representative structural and functional information for the protein domain family that includes AF2095.
This document includes the supplementary material originally published online only.


Comments
Published in Journal of Biomolecular NMR 30 (2004), pp. 107–108. http://www.springerlink.com/content/1573-5001/ Copyright © 2004 Kluwer Academic Publishers. Used by permission.
This document includes the supplementary material originally published online only.