Plant Pathology, Department of
Department of Plant Pathology: Faculty Publications
Document Type
Article
Date of this Version
2007
Citation
Jo, Y.-K., Wang, G.-L., and Boehm, M. J. 2007. Expression analysis of rice defense-related genes in turfgrass in response to Magnaporthe grisea. Phytopathology 97:170-178.
Abstract
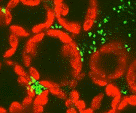
Magnaporthe grisea (anamorph = Pyricularia grisea) causes blast on rice (Oryza sativa) and gray leaf spot on turfgrass. Gray leaf spot is a serious disease on St. Augustinegrass (Stenotaphrum secundatum), perennial ryegrass (Lolium perenne), and tall fescue (Festuca arundinacea). Virulence assays performed in this study revealed that M. grisea collected from rice could also cause disease on St. Augustinegrass and tall fescue. One rice isolate, Che86061, caused similar disease reactions on susceptible cultivars of rice and St. Augustinegrass and an incompatible interaction on resistant cultivars of both species. To explore whether similar defense-related genes are expressed in rice and St. Augustinegrass, a rice cDNA library was screened using pooled cDNAs derived from M. grisea infected St. Augustinegrass. Thirty rice EST (expressed sequence tag) clones showing differential expression in St. Augustinegrass following M. grisea inoculation were identified and classified into six putative functional groups. Northern blot analyses of seven EST clones that collectively represented each putative functional group confirmed that the expression of five out of seven EST clones was similar in both rice and St. Augustinegrass. This study represents one of the first attempts to use a broad-scale genomic approach and resources of a model monocot system to study defense gene expression in St. Augustinegrass following M. grisea infection.


Comments
Open Access licensed.