Chemistry, Department of: Faculty Series
David Hage Publications
Document Type
Article
Date of this Version
9-1-2010
Citation
Published in final edited form as: Anal Biochem. 2010 September 1; 404(1): 106–108. doi:10.1016/j.ab.2010.05.004. Version presented here is from NIH PubMed Central
Abstract
Affinity chromatography uses a biologically-related binding agent as a stationary phase to undergo specific interactions with a target analyte (1–3). The selectivity of this approach is dependent on the way in which the binding agent is attached to the support (4). It is common for a binding agent such as a protein to be covalently immobilized through amine, sulfhydryl, carboxyl or carbonyl groups (4–6). However, covalent immobilization can produce multisite attachment or improper orientation of the binding agent, which may cause an apparent change in this agent’s activity. These issues can be minimized in some cases by coupling the binding agent to the support through functional groups that occur in only a few locations in its structure, such as the use of the carbohydrate chains in antibodies or α1-acid glycoprotein (AGP) for their site-selective attachment to hydrazide-activated supports (4,6).
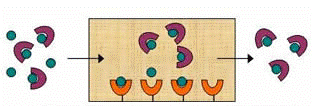
Another approach to avoid these immobilization effects is to use the physical entrapment (or encapsulation) of the binding agent within a support. This approach has been of interest for many years in work with materials such as sol gels and has generally been based on the physical containment of binding agents in a highly cross-linked polymer network (7–9). However, past approaches have used materials with pressure or flow rate restrictions that are not easily amenable for use with HPLC. Also, a general entrapment approach is still needed that can be used with common HPLC supports such as porous silica.


Comments
Copyright Elsevier Inc. Used by permission.